fmicb-11-01171 June 1, 2020 Time: 18:7 # 1
HYPOTHESIS AND THEORY
published: 03 June 2020
doi: 10.3389/fmicb.2020.01171
Edited by:
Andrzej Gorski,
Polish Academy of Sciences, Poland
Reviewed by:
Stephen Tobias Abedon,
The Ohio State University,
United States
Ronen Nissan Hazan,
The Hebrew University of Jerusalem,
Israel
*Correspondence:
Jean-Paul Pirnay
Specialty section:
This article was submitted to
Antimicrobials, Resistance
and Chemotherapy,
a section of the journal
Frontiers in Microbiology
Received: 27 February 2020
Accepted: 07 May 2020
Published: 03 June 2020
Citation:
Pirnay J-P (2020) Phage Therapy
in the Year 2035.
Front. Microbiol. 11:1171.
doi: 10.3389/fmicb.2020.01171
Phage Therapy in the Year 2035
Jean-Paul Pirnay
*
Laboratory for Molecular and Cellular Technology, Queen Astrid Military Hospital, Brussels, Belgium
The emergence of multidrug resistant bacteria in both community- and hospital-
acquired infections is recognized as a major public health threat. Phage therapy is
increasingly mediatized and researched as an additional tool for combatting antibiotic
resistant infections. However, phages exhibit a number of properties that differ from
antibiotics and hamper their development as pharmaceutical products and their
application in therapy. This paper advocates a paradigm shift in the development and
application of infectious disease therapeutics to cater for personalized phage therapy,
which could be realized by the year 2035. More specifically, it presents a sustainable
and ethical supply chain of instant synthetic phages, based on a community effort,
supported and steered by public health organizations, and managed by a platform
combining Artificial Intelligence (AI) and Distributed Ledger (DL) Technology.
Keywords: infectious diseases, antibiotic resistance, antimicrobial resistance, phage therapy, synthetic biology,
artificial intelligence, machine learning, distributed ledger technology
PREFACE
This paper offers a personal vision of what might be needed for phage therapy to finally break
through as a mainstream antibacterial tool. It is influenced by historical and recent failures and
uncertainties in the phage therapy field and aims at finding solutions based on future and emerging
technologies that are supposed to model the science and society of tomorrow.
PHAGE THERAPY
Bacteriophages (phages) are the viruses of bacteria. Since time immemorial they have controlled
the growth and spread of their bacterial hosts. Bacterial viruses are the most ubiquitous lifelike
entities in our biosphere. There are an estimated 10 million-fold more viruses in the oceans than
there are stars in the universe and if all the phages on Earth were stacked on top of another, this
tower would stretch further than the nearest 60 galaxies (Suttle, 2013). They can easily be found
wherever bacteria thrive: in sewers, rivers, or patients’ urine and stool. Phages of human bacterial
pathogens are most often composed of an icosahedral head, a sphere with 20 flat faces made of
proteins and containing a nucleic acid genome, to which a protein tail is attached. When a strictly
lytic phage adheres with its tail fibers and spikes to the surface of its target bacterium, the syringe-
like tail sheath contracts and the tail core is driven through the bacterial cell wall, injecting the
phage genome into the periplasm of the bacterial cell. Immediately, the bacterial DNA and protein
synthesis machinery is hijacked to build copies of the phage. Some phages cut the bacterial DNA
into pieces. After a latent period of minutes to hours, the newly formed phages burst out of their
bacterial hosts, which are killed in the process. The phage progeny, which can run into the hundreds
per bacterium, then go off to find new host bacteria to infect. As such, phages can be considered
Frontiers in Microbiology | www.frontiersin.org 1 June 2020 | Volume 11 | Article 1171

fmicb-11-01171 June 1, 2020 Time: 18:7 # 2
Pirnay Phage Therapy in 2035
as self-replicating antimicrobials. Importantly, phages have
evolved to only infect certain target bacteria and are harmless to
mammalian cells.
Early evidence of viral-like agents with antibacterial activity
was reported by the English bacteriologist Frederick Twort,
and by the French-Canadian microbiologist Felix d’Hérelle
in 1915 and 1917, respectively (Sulakvelidze et al., 2001). In
1919, d’Hérelle exploited for the first time the therapeutic
potential of phages when he used them to cure a boy suffering
from dysentery in Paris. Phage therapy was immediately
recognized as a therapeutic approach to treat bacterial infections
and commercialization of phage therapy preparations was
undertaken by several companies, such as L’Oréal in Europe and
Eli Lilly Company in the United States (Sulakvelidze et al., 2001).
In 1923, the Georgian microbiologist Giorgi Eliava founded the
Eliava Institute in Tbilisi, Georgia, devoted to phage therapy
research. It was the start of extensive phage therapy research
and development in the former Soviet-Union. However, early
uses of phage therapy were often unreliable and research into
antibiotics had also been ongoing. The successful use of penicillin
during the Second World War and its subsequent worldwide
marketing led Western scientists to lose interest in phage
therapy. Soviet researchers, in contrast, continued to develop
phage therapy and to publish their results, but due to the
Iron Curtain their knowledge and experience did not spread
across the world (Sulakvelidze et al., 2001). At the dawn of the
third millennium, the increasing health burden of infections
with antibiotic resistant bacteria (Cassini et al., 2019) incited
a renewed worldwide interest in phage therapy as a viable
additional tool to the clinical management of bacterial infections
(Thiel, 2004). All over the world, phage therapy centers are being
set up, following in the footsteps of the Eliava Institute and the
phage therapy unit at the Hirszfeld Institute in Wrocław, Poland
(Miêdzybrodzki et al., 2012).
THE YEAR 2035
Fast forward to future Earth of 2035, a gloomy world
characterized by human overpopulation, major ecosystem
disruptions, global warming, and xenophobia.
While soaking in his bath, Dr. John Iverian, a retired
microbiologist, suddenly felt an extremely painful sting in the
back of his neck, followed by a sound like a small plane’s
propeller. He screened the environment and in the corner of
his eye he saw a weird large insect with long creepy legs and
antennas sitting on the wall next to his designer bathtub. Osuri,
the home management system of Iverian’s loft in the center of
Antwerp, identified the insect as the brown marmorated stink
bug Halyomorpha halys, which had spread across the world.
Osuri’s report, projected on one of the bathroom’s video screens,
mentioned that people, who were bitten, initially experienced a
small red sore in the bite area of their skin. When left unattended,
the bite wound would swell and produce puss. Tired and muzzy,
non-chalant Iverian stepped out of his bath and went to bed.
He had decided not to perform the elaborate wound treatment
procedure, which had strongly been advised by Osuri. Early next
morning, however, the bite had turned into a necrotic wound
showing clear signs of infection.
Anxiously, Iverian activated his Phage-BEAM device. BEAM
stood for “Bedside Energized Anti-Microbial.” The device had the
size and shape of a shoebox, but with a more elegant and polished
look. The name of the device and its manufacturer were designed
in colorful letters on the side of the seamless white enameled
box. Iverian removed a swab from its sterile packaging and gently
passed it over the entire area of the wound, making sure that
the wound exudate thoroughly wetted the cotton wool tip of the
swab. When the swab approached the “insert sample” area of the
box, a tiny door opened as if magically, freeing a 10-inch high
hologram of a lab technician, named Marcia. She showed Iverian
where to dock the sample. Marcia was developed to guide the
clients through the test procedure. “For best results, please insert
a new phage bio-ink cartridge, Dr. Iverian,” Marcia said. Just as
it used to be for yesteryear’s 2D printers, the cost of the bio-ink
cartridges was almost as high as the cost of the Phage-BEAM
device itself. According to “Business Insider,” phage bio-ink was
the second most expensive liquid on Earth, behind Chanel No.
8. Luckily, as one of the inventors, Iverian had obtained the
right to always have the most recent version of this device at his
disposition, including a continuous supply of reagents, for free.
Iverian knew perfectly how the device worked, so he did not
need Marcia’s help. First, DNA was extracted from the swab tip
and the metagenome—all the genetic material present in the
sample, including the infecting bacteria—was determined. Next,
these genetic data were sent to a secured “Phage XChange” server
where a complex AI-driven algorithm predicted the genome
sequence of the phage that was most likely to lyze the infecting
bacteria identified in the metagenome and was supposed to elicit
the weakest immune reaction in the patient. The phage genome
data were sent to the Phage-BEAM device, which first synthesized
the phage genome and then the phage, using a proprietary
bacterium-free phage production system.
Within 1 h after sampling, the device would produce a ready-
to-use therapeutic phage product. Results of the step-by-step
procedure would be transmitted to the enormous home video
screen in Iverian’s living room. Sitting in his LC2 armchair,
listening to Mozart’s Great Mass in C minor, Iverian anxiously
waited for the results to come in. He had a bad feeling about
this. The result sent shivers down Iverian’s spine. Bacterial
pathogen identified: Streptococcus pyogenes strain FE-2033! Osuri
immediately activated the infection alert protocol, sending a
message to the World Center for Disease Control and projected
worrisome background information on the lethal flesh eating
bacterial strain, which was considered an imminent threat to
public health since 2033. For a moment, Iverian considered
excising the infected wound and some surrounding healthy tissue
with a kitchen knife, but he calmed down and decided to wait
and apply the imminent Phage-BEAM product. An hour later,
the Phage-BEAM device had produced synthetic phages. These
phages were then mixed with the isolated bacteria, in a validation
module, to test their in vitro efficacy. Fifteen minutes later, the
green light was given for Iverian to commence treatment. Iverian
applied the phages in a slow release hydrogel-based wound
dressing, which had first been mixed with the concentrated
Frontiers in Microbiology | www.frontiersin.org 2 June 2020 | Volume 11 | Article 1171

fmicb-11-01171 June 1, 2020 Time: 18:7 # 3
Pirnay Phage Therapy in 2035
phage suspension produced by the Phage-BEAM device, and
also contained synergistic antibiotics. The hydrogel temporarily
relieved the pain, which further calmed him. Iverian repeated
application of the phage and antibiotic-loaded hydrogel once a
day. Wound infection improved within 24 h and after 7 days the
wound was almost completely healed. Iverian’s potentially life-
threatening infection was successfully treated, timely and without
leaving his home. But, for many previous decades, it had not
been certain that phage therapy would break through to become
a broadly applied and clinically useful antibacterial tool. The
medical world had taken a while to realize that phage therapy
did not need to be identical to antibiotic therapy, and this mainly
because of the peculiarities of the active agents, the phages.
SOME RELEVANT PECULIARITIES OF
PHAGES
Phages exhibit a number of properties that differ from antibiotics
and hampered their development as pharmaceutical products
and application in therapy. First, they tend to be specific
about which bacteria they infect. They will at best target a
considerable part of one single bacterial species, but at worst
they will infect only a small number of strains within one
species. Therapeutic phages can thus be selected to mainly kill
one bacterial species, or a clinically relevant subgroup thereof,
and spare the patient’s beneficial bacteria (e.g., the gut, skin, or
oral commensal flora). Most routinely employed antibiotics, in
contrast, have a broad spectrum of activity, which can cause
“collateral damage” to the patient’s commensal microbiomes,
which in turn can result in adverse effects such as the selection
of antibiotic resistant bacterial species (e.g., Clostridium difficile)
or antibiotic-associated diarrhea (Jernberg et al., 2010). The
drawback of phage specificity is that the infecting bacteria need to
be identified before starting phage therapy. In empirical antibiotic
therapy, in contrast, broad-spectrum antibiotic cocktails that
affect a multitude of Gram-positive and Gram-negative bacteria,
and diverse fungi are typically used. When more information is
known (e.g., from bacterial culture), treatment may consist of
narrow-spectrum antibiotics, which more specifically target the
bacteria or fungi identified to be causing disease.
Second, bacteria and phages are involved in a host–
parasite relationship. Strictly lytic phages are ubiquitous in the
environment and require the death of their bacterial host to
complete their life cycle. Without hosts, phages cannot exist.
Phages impose selection for resistant hosts, which in turn
impose selection for effective phages. This results in what is
called “antagonistic coevolution,” an arms race between bacteria
and phages, characterized by reciprocal evolution of bacterial
resistance and phage infectivity (Buckling and Rainey, 2002).
Just as with most antimicrobials, bacteria will thus also become
resistant to phages (Luria and Delbrück, 1943; Schooley et al.,
2017), but, in contrast to static antibiotics, phages have the
capacity to overcome bacterial resistance (Buckling and Rainey,
2002). There are nevertheless indications that bacteria and phages
will not indefinitely increase their respective resistance and
infectivity (Fortuna et al., 2019).
PHAGE THERAPY APPROACHES
At the time of the phage therapy revival in the early 2000s,
two distinct phage therapy approaches had been developed
(Pirnay et al., 2011). In what could be called the one-size-fits-all
approach, defined broad-spectrum phage cocktails, which were
supposed to target the majority of bacteria suspected to cause
certain infectious diseases, were applied. These predefined broad-
spectrum phage cocktails were developed, produced, and tested
within the current pharmaco-economic models, which had been
designed to cater for “static” drugs such as antibiotics. However,
truly broad-spectrum phage cocktails, active against most Gram-
positive and/or Gram-negative bacteria commonly encountered
in infectious diseases needed to contain large amounts of phages
and turned out to be very difficult to develop. It was feasible
to develop narrower spectrum phage cocktails, active against
one or a few bacterial species, to be used in certain indications
and minding that the infecting bacterial species were known
in advance. For some bacterial species, such as Staphylococcus
aureus, phages showing an exceptionally broad host range had
been isolated and characterized (Vandersteegen et al., 2011). In
PhagoBurn, a randomized controlled trial, success in achieving
sustained reduction in Pseudomonas aeruginosa burdens in burn
wounds was linked to initial susceptibility to the phage cocktail
(Jault et al., 2019). However, one-third of the included patients
were shown to harbor pre-existing P. aeruginosa strains resistant
to the phage cocktail, which consisted of no less than 12
lytic P. aeruginosa phages. In addition, phage cocktails that
were initially designed to be effective needed to be regularly
updated (e.g., supplemented with new phages) in response
to the emergence of phage resistance or the involvement of
newly circulating clinically relevant strains. Finally, it was not
known if confronting bacteria with high concentrations of
fixed phage cocktails would cause the emergence, spread and
persistence of bacterial phage resistance in hospitals and in the
environment, similar to what had happened upon the massive
use of antibiotics.
In personalized phage therapy concepts, one or more phages
were selected from a phage bank, or from the environment,
and possibly adapted (in vitro selection of phage mutants
exhibiting increased infectivity) to more efficiently infect the
bacteria isolated from the patient’s infection site (Friman et al.,
2016). Some phage therapy centers set up and maintained large
therapeutic phage banks, which were regularly updated with
new phages, widening and adapting the host range of the bank
to the ever-changing bacterial populations. Personalized phage
therapy approaches were potentially more sustainable, as only
the infecting bacterium is targeted, resulting in less selection
pressure toward bacterial phage resistance. However, they were
also more elaborate and logistically complex than one-size-fits-
all approaches, with bacterial strains and matching phages being
sent all over the world (Figure 1). Moreover, precision medicine
concepts were, in general, not compatible with most medicinal
product (drugs in the United States) development and licensing
pathways, which required several years and millions of euros
(dollars) to complete, and this for every phage in the bank
(Verbeken et al., 2012).
Frontiers in Microbiology | www.frontiersin.org 3 June 2020 | Volume 11 | Article 1171

fmicb-11-01171 June 1, 2020 Time: 18:7 # 4
Pirnay Phage Therapy in 2035
FIGURE 1 | International transfers of phages from (red arrows) and to (blue arrows) the Queen Astrid Military Hospital (QAMH) in Brussels in view of clinical
applications over the period 2015–2020. On the national level, phages were dispatched from the QAMH to five university hospitals (not shown). In addition, the
selection of matching phages often encompassed the transfer of the patients’ bacterial isolates, and five international patients (two from France, two from the
Netherlands, and one from Tunisia) were transferred to Brussels for phage therapy.
ENTER SYNTHETIC BIOLOGY
With the onset of the third millennium, synthetic biology
approaches had been increasingly developed to reduce the
specificity of phages and the emergence of bacterial phage
resistance (e.g., structure-guided design) (Pires et al., 2016;
Dunne et al., 2019). For instance, yeast-based platforms for phage
tail fiber protein switching were elaborated to engineer hybrid
phages with more predictable and extended host range (Ando
et al., 2015; Yosef et al., 2017) and genetic engineering strategies
(e.g., CRISPR-Cas editing tools) were developed to address other
aspects such as negative patient-phage interactions (e.g., anti-
phage immune response) (Brown et al., 2017), the potential
emergence and spread of bacterial phage resistance mechanisms,
and the release of harmful bacterial contents such as endotoxins
(Hwang et al., 2018). Synthetic phage genomes needed to be
rebooted to produce phage offspring (Barbu et al., 2016; Pires
et al., 2016), through transformation of Escherichia coli or Listeria
monocytogenes L-forms (Kilcher et al., 2018), or using cell-free
transcription-translation (TXTL) systems (Rustad et al., 2018).
Western regulatory frameworks had gradually started to cater
for precision and personalized phage therapy approaches using
naturally occurring phages (Pirnay et al., 2018), engineered
phages (Dedrick et al., 2019), and synthetic phages.
The development of ad hoc and on-site therapeutic phage
production devices, such as Phage-BEAM, did not run smoothly,
Iverian recalled. To start with, it required artificial intelligence
(AI)-based in silico phage matching and design. Deep learning
(Martorell-Marugán et al., 2019), a subset of Machine Learning,
was chosen to search for links between bacterial genomes and
infecting phage genomes, because it was easier to scale to bigger
number of samples. For instance, deep learning methods did
not require so-called feature extraction, which would require
gene/protein level annotation of phage and bacterial genomes
and would limit predictions to certain known relationships
between bacterial and phage features, such as phage tail fiber
structures binding to specific bacterial cell wall receptors. As a
down side, it needed to be powered by a continuous supply of
massive amounts of data, linking lytic phage genomes to host
bacterial genomes, and that’s where the shoe pinched. Whole
genome sequencing had slowly percolated into the practice
of clinical microbiology (American Academy of Microbiology,
2016), but research institutes and pharmaceutical companies
were not keen to submit their data to a single centralized
database, and no investors were found willing and able to
acquire the available data and/or to produce sufficient amounts
of new data. A second obstacle that had to be overcome was
the unavailability of quick, reliable, and affordable synthesis of
Frontiers in Microbiology | www.frontiersin.org 4 June 2020 | Volume 11 | Article 1171
fmicb-11-01171 June 1, 2020 Time: 18:7 # 5
Pirnay Phage Therapy in 2035
large DNA molecules. Initial DNA synthesis techniques were
based on organic chemistry and produced relatively small DNA
molecules. The de novo synthesis of phage genomes required
assembling several genome fragments (Barbu et al., 2016; Pires
et al., 2016; Lemire et al., 2018) in the yeast Saccharomyces
cerevisiae, using yeast artificial chromosomes (Ando et al., 2015),
or chemical assembly (Gibson et al., 2009). The development
of a new technique to synthesize DNA, based on DNA-
synthesizing enzymes found in cells of the immune system
(Palluk et al., 2018), facilitated phage genome synthesis. Finally,
some hurdles had to be overcome to develop generic cell-free
phage production systems able to produce phages in high titers
and exhibiting the same levels of bacterial infectivity as their
natural analogs.
THE BREAKTHROUGH
The major problem was that it turned out to be very difficult
to collect the massive amounts of linked phage and bacterial
genome sequences necessary for the deep learning AI algorithms
to predict and/or design phage sequences with a therapeutically
acceptable level of accuracy. Iverian remembered that the real
breakthrough came when the not-for-profit organization “Phage
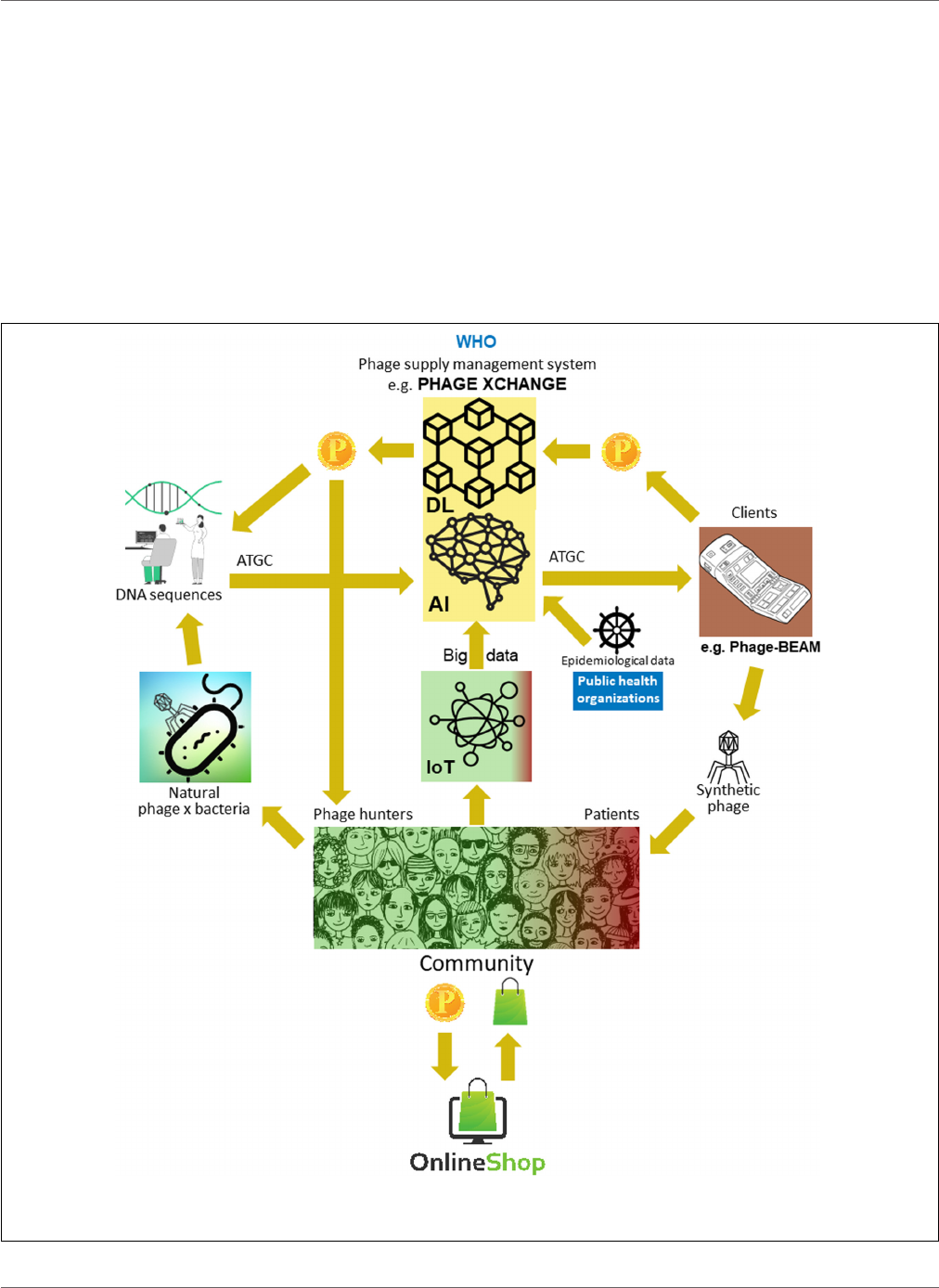
FIGURE 2 | Vision of how the phage supply chain might be organized in 2035. AI, artificial intelligence; ATGC, DNA sequence; BEAM, bedside energized
anti-microbial; DL, distributed ledger; IoT, Internet of Things; P, PhageCoin; WHO, World Health Organization.
Frontiers in Microbiology | www.frontiersin.org 5 June 2020 | Volume 11 | Article 1171

fmicb-11-01171 June 1, 2020 Time: 18:7 # 6
Pirnay Phage Therapy in 2035
XChange” launched its global phage governance platform of the
same name to create an efficient, standardized, sustainable, and
ethical phage supply chain (Figure 2). Phage XChange mainly
consisted of an AI module and a Distributed Ledger (DL)
(Thiebes et al., 2020). The platform’s AI module analyzed linked
phage and bacterial genomes to predict and design potent phages
for clients. It also predicted which bacterial pathogens needed the
most urgent attention, based on the Internet of Things (IoT) and
Big Data and information provided by international public health
organizations, such as the World Health Organization (WHO)
and national Centers for Disease Control. These data steered
the system toward the isolation and characterization of the most
urgently needed phages.
The platform’s DL module ensured a sufficient, qualitative,
and recorded input of linked phage/bacterial genome sequences
to the AI module and ditto supply of phage sequences to
clients, in compliance with the provisions of the Nagoya protocol
(Expert round table on acceptance and re-implementation of
bacteriophage therapy et al., 2018). The DL immutably recorded
all stakeholder (e.g., suppliers of data, sequencing services, and
clients), transaction, and contract details. It also recorded the
exact quality, specifications, and weight of the supplied material.
An algorithm determined the non-redundancy and estimated
the weight (e.g., the virulence and host range of the phages)
and desirability of the submitted material. Phages targeting
emerging bacterial pathogens were of course most wanted.
Most patent issues were obviated. The DL acted as a payment
ledger to assure that all parties were paid timely and fairly.
A number of PhageCoins (the platform’s crypto currency) were
attributed to the suppliers in relation to the quality, weight, and
desirability of the supplied material. Clients extracting prediction
results (phage sequences) through the DL paid an amount of
PhageCoins, proportionate to the estimated value of the phages.
These PhageCoins were used to maintain the DL, to assure a
sufficient and continuous inflow of material, and to expand phage
virulence and host range data (matching phages to bacteria). An
additional injection of funds and incentive to supply material
was found in producers and suppliers of all kinds of goods.
With the instantly earned PhageCoins, phage suppliers could
buy online all kinds of products at strongly reduced prices,
from laboratory- and school equipment to sports items. These
goods were provided through corporate sponsorship. Several
established companies sponsored PhageXchange in exchange of
tax reductions, publicity, and the image of a socially responsible
brand. The weight of the supplied material, and thus also
its value, were initially undervalued, but were re-evaluated at
regular intervals (iteration) and suppliers were attributed more
PhageCoins when warranted. Even though useful from the
moment it was introduced, the platform only became really
successful when it was put under the protection of the WHO,
in analogy to the worldwide system of traceability, transparency,
vigilance, and surveillance of Medicinal Products of Human
Origin (Warwick et al., 2013). A formal agreement between Phage
XChange and the WHO increased international confidence in
the long-term sustainability of the platform and protection
from unethical commercial exploitation. The search for potent
therapeutic phages soon became a community effort aimed
at solving the antibiotic resistance crisis, with independent
“phage hunters,” schools, scout groups, villages on the banks
of the Amazon River, etc., isolating and submitting phages to
Phage XChange, in exchange of PhageCoins. At the margins of
this, various companies and institutions developed peripheral
equipment and services, such as phage isolation kits and
sequencing and phage synthesizing platforms (e.g., the Phage-
BEAM device). In anticipation of these devices, intermediary
solutions were offered, whereby the phages themselves were
obtained through the DL.
EPILOG
This view on the future of phage therapy provides an optimistic
ending to the antibiotic resistance crisis. The ad hoc and
on site production of synthetic phages, linked to a global,
community-based, phage management system, turned out to
be a welcome and affordable (for social security systems)
extra weapon in the fight against antibiotic resistant bacterial
infections. However, it was not a magic bullet; it was a synergistic
supplement to established antimicrobials. The instant and cell-
free production of synthetic phages, whether designed or not, had
considerable advantages over classically produced (in bacterial
hosts) natural phages:
(i) There was no need to maintain physical therapeutic phage
banks and to dispatch the patient’s bacterial isolates and the
matching therapeutic phages all over the world.
(ii) Synthetic phages against bacteria causing eminent public
health threats, such as the 2011 E. coli O104:H4 outbreak in
Germany (Merabishvili et al., 2012), or bacteria (suspected
to be) used for bioterrorism (Joñczyk-Matysiak et al., 2014)
could be timely produced on site.
(iii) Phages against bacteria causing potentially lethal diseases,
for which no non-lethal production host strains were
available and whose propagation used to require biosafety
level-3 (bsl-3) bio-containment precautions, could be
synthesized in bsl-1 conditions.
(iv) When no phages could be isolated from sampling sites,
for instance, because the bacterial host strains used in the
isolation techniques were not susceptible to the desired
phages, (predicted) phage genomic sequences, extracted
from metagenomic data (Reyes et al., 2010; Amgarten et al.,
2018), could be used to produce synthetic phages.
(v) Synthetic phage preparations contained no (or smaller
amounts of) molecules that could have a negative impact
on patients (e.g., endotoxins).
(vi) Devices were adapted to produce synthetic phages
during extended space travel and space colonization
(Taylor and Sommer, 2005).
There is little chance that these predictions will come true.
It is probably too shortsighted to think that a community-based
effort, supported by public health organizations and managed by
a global, sustainable and ethical platform, could be at the heart
of a solution to the current worldwide antibiotic resistance crisis.
Frontiers in Microbiology | www.frontiersin.org 6 June 2020 | Volume 11 | Article 1171

fmicb-11-01171 June 1, 2020 Time: 18:7 # 7
Pirnay Phage Therapy in 2035
Some parts of the proposed system, such as cell free production
of synthetic phages using a bedside device, have a reasonable
chance of being realized, while other elements, such as
corporate sponsorship, will likely remain limited to the
realm of science fiction. You may say that I’m a dreamer,
so feel free to wake me up in 2035 to confront me
with reality!
AUTHOR CONTRIBUTIONS
J-PP conceived the vision and drafted the manuscript.
FUNDING
Publication costs were covered by “Société Scientifique du
Service Médical Militaire – Wetenschappelijke Vereniging van de
Militaire Medische Dienst”.
ACKNOWLEDGMENTS
The personal vision, or dream, developed in this manuscript
came about as a result of interactions with many fellow
researchers over the past 15 years. It is impossible to name
them all, but it would not be fair to take all the credits alone.
Therefore, I decided to acknowledge some of them here (in
alphabetical order), with the risk—or better, the certainty—
of forgetting some important influencers: Joana Azeredo, Nata
Bakuradze, Bob Blasdel, Dimitri Boeckaerts, Angus Buckling,
Yves Briers, Pieter-Jan Ceyssens, Nina Chanishvili, Laurent
Debarbieux, Sarah Djebara, Dorien Dams, Daniel De Vos,
Quirin Emslander, Alan Fauconnier, Ville Friman, Andrzej
Górski, Téa Glonti, Nino Grdzelishvili, Serge Jennes, Elene
Kakabadze, Betty Kutter, Rob Lavigne, Cédric Lood, Alice
Maestri, Khatuna Makalatia, Maya Merabishvili, Tobi Nagel,
Thomas Rose, Patrick Soentjens, Michiel Stock, Rüdiger Trojok,
An Van den Bossche, Mario Vaneechoutte, Gilbert Verbeken,
and Kilian Vogele.
REFERENCES
American Academy of Microbiology (2016). Applications of Clinical Microbial
Next-Generation Sequencing: Report on an American Academy of Microbiology
Colloquium Held in Washington, DC, in April 2015. Washington, DC: American
Society for Microbiology.
Amgarten, D., Braga, L. P. P., da Silva, A. M., and Setubal, J. C. (2018). MARVEL,
a tool for prediction of bacteriophage sequences in metagenomic bins. Front.
Genet. 9:304. doi: 10.3389/fgene.2018.00304
Ando, H., Lemire, S., Pires, D. P., and Lu, T. K. (2015). Engineering modular
viral scaffolds for targeted bacterial population editing. Cell. Syst. 1, 187–196.
doi: 10.1016/j.cels.2015.08.013
Barbu, E. M., Cady, K. C., and Hubby, B. (2016). Phage therapy in the era of
synthetic biology. Cold Spring Harb. Perspect. Biol. 8:a023879. doi: 10.1101/
cshperspect.a023879
Brown, R., Lengeling, A., and Wang, B. (2017). Phage engineering: how advances
in molecular biology and synthetic biology are being utilized to enhance the
therapeutic potential of bacteriophages. Quant. Biol 5, 42–54.
Buckling, A., and Rainey, P. B. (2002). Antagonistic coevolution between a
bacterium and a bacteriophage. Proc. Biol. Sci. 269, 931–936. doi: 10.1098/rspb.
2001.1945
Cassini, A., Högberg, L. D., Plachouras, D., Quattrocchi, A., Hoxha, A., Simonsen,
G. S., et al. (2019). Attributable deaths and disability-adjusted life-years caused
by infections with antibiotic-resistant bacteria in the EU and the European
Economic Area in 2015: a population-level modelling analysis. Lancet Infect.
Dis. 19, 56–66. doi: 10.1016/S1473-3099(18)30605-4
Dedrick, R. M., Guerrero-Bustamante, C. A., Garlena, R. A., Russell, D. A., Ford,
K., Harris, K., et al. (2019). Engineered bacteriophages for treatment of a patient
with a disseminated drug-resistant Mycobacterium abscessus. Nat. Med. 25,
730–733. doi: 10.1038/s41591-019-0437-z
Dunne, M., Rupf, B., Tala, M., Qabrati, X., Ernst, P., Shen, Y., et al. (2019).
Reprogramming bacteriophage host range through structure-guided design of
chimeric receptor binding proteins. Cell Rep. 29, 1336.e4–1350.e4. doi: 10.1016/
j.celrep.2019.09.062
Expert round table on acceptance and re-implementation of bacteriophage therapy,
Sybesma, W., Rohde, C., Bardy, P., Pirnay, J.-P., Cooper, I., et al. (2018). Silk
route to the acceptance and re-implementation of bacteriophage therapy-part
II. Antibiotics 7:E35. doi: 10.3390/antibiotics7020035
Fortuna, M. A., Barbour, M. A., Zaman, L., Hall, A. R., Buckling, A., and
Bascompte, J. (2019). Coevolutionary dynamics shape the structure of bacteria-
phage infection networks. Evolution 73, 1001–1011. doi: 10.1111/evo.13731
Friman, V. P., Soanes-Brown, D., Sierocinski, P., Molin, S., Johansen, H. K.,
Merabishvili, M., et al. (2016). Pre-adapting parasitic phages to a pathogen
leads to increased pathogen clearance and lowered resistance evolution with
Pseudomonas aeruginosa cystic fibrosis bacterial isolates. J. Evol. Biol. 29,
188–198. doi: 10.1111/jeb.12774
Gibson, D. G., Young, L., Chuang, R. Y., Venter, J. C., Hutchison, C. A. III,
and Smith, H. O. (2009). Enzymatic assembly of DNA molecules up to
several hundred kilobases. Nat. Methods 6, 343–345. doi: 10.1038/nmeth.
1318
Hwang, I. Y., Lee, H. L., Huang, J. G., Lim, Y. Y., Yew, W. S., Lee, Y. S., et al. (2018).
Engineering microbes for targeted strikes against human pathogens. Cell Mol.
Life. Sci. 75, 2719–2733. doi: 10.1007/s00018-018-2827-7
Jault, P., Leclerc, T., Jennes, S., Pirnay, J. P., Que, Y.-A., Resch, G., et al. (2019).
Efficacy and tolerability of a cocktail of bacteriophages to treat burn wounds
infected by Pseudomonas aeruginosa (PhagoBurn): a randomised, controlled,
double-blind phase 1/2 trial. Lancet Infect. Dis. 19, 35–45. doi: 10.1016/S1473-
3099(18)30482-1
Jernberg, C., Löfmark, S., Edlund, C., and Jansson, J. K. (2010). Long-term impacts
of antibiotic exposure on the human intestinal microbiota. Microbiology 156,
3216–3223. doi: 10.1099/mic.0.040618-0
Joñczyk-Matysiak, E., Kłak, M., Weber-Da¸browska, B., Borysowski, J., and Górski,
A. (2014). Possible use of bacteriophages active against Bacillus anthracis and
other B. cereus group members in the face of a bioterrorism threat. Biomed. Res.
Int. 2014:735413. doi: 10.1155/2014/735413
Kilcher, S., Studer, P., Muessner, C., Klumpp, J., and Loessner, M. J. (2018).
Cross-genus rebooting of custom-made, synthetic bacteriophage genomes in
L-form bacteria. Proc. Natl. Acad. Sci. U.S.A. 115, 567–572. doi: 10.1073/pnas.
1714658115
Lemire, S., Yehl, K. M., and Lu, T. K. (2018). Phage-based applications in synthetic
biology. Annu. Rev. Virol. 5, 453–476. doi: 10.1146/annurev-virology-092917-
043544
Luria, S. E., and Delbrück, M. (1943). Mutations of bacteria from virus sensitivity
to virus resistance. Genetics 28, 491–511.
Martorell-Marugán, J., Tabik, S., Benhammou, Y., del Val, C., Zwir, I., Herrera, F.,
et al. (2019). “Deep learning in omics data analysis and precision medicine,”
in Computational Biology, ed. H. Husi (Brisbane: Codon Publications), 37–53.
doi: 10.15586/computationalbiology.2019.ch3
Merabishvili, M., De Vos, D., Verbeken, G., Kropinski, A. M., Vandenheuvel,
D., Lavigne, R., et al. (2012). Selection and characterization of a candidate
therapeutic bacteriophage that lyses the Escherichia coli O104:H4 strain from
the 2011 outbreak in Germany. PLoS One 7:e52709. doi: 10.1371/journal.pone.
0052709
Miêdzybrodzki, R., Borysowski, J., Weber-Da¸browska, B., Fortuna, W., Letkiewicz,
S., Szufnarowski, K., et al. (2012). Clinical aspects of phage therapy. Adv. Virus
Res. 83, 73–121. doi: 10.1016/B978-0-12-394438-2.00003-7
Frontiers in Microbiology | www.frontiersin.org 7 June 2020 | Volume 11 | Article 1171

fmicb-11-01171 June 1, 2020 Time: 18:7 # 8
Pirnay Phage Therapy in 2035
Palluk, S., Arlow, D. H., de Rond, T., Barthel, S., Kang, J. S., Bector, R., et al.
(2018). De novo DNA synthesis using polymerase-nucleotide conjugates. Nat.
Biotechnol. 36, 645–650. doi: 10.1038/nbt.4173
Pires, D. P., Cleto, S., Sillankorva, S., Azeredo, J., and Lu, T. K. (2016).
Genetically engineered phages: a review of advances over the last
decade. Microbiol. Mol. Biol. Rev. 80, 523–543. doi: 10.1128/MMBR.00
069-15
Pirnay, J. P., De Vos, D., Verbeken, G., Merabishvili, M., Chanishvili, N.,
Vaneechoutte, M., et al. (2011). The phage therapy paradigm: prêt-à-
porter or sur-mesure? Pharm. Res. 28, 934–937. doi: 10.1007/s11095-010-
0313-5
Pirnay, J. P., Verbeken, G., Ceyssens, P. J., Huys, I., De Vos, D., Ameloot,
C., et al. (2018). The magistral phage. Viruses 10:E64. doi: 10.3390/v1002
0064
Reyes, A., Haynes, M., Hanson, N., Angly, F. E., Heath, A. C., Rohwer, F., et al.
(2010). Viruses in the fecal microbiota of monozygotic twins and their mothers.
Nature 466, 334–338. doi: 10.1038/nature09199
Rustad, M., Eastlund, A., Jardine, P., and Noireaux, V. (2018). Cell-free TXTL
synthesis of infectious bacteriophage T4 in a single test tube reaction. Synth.
Biol. 3:ysy002.
Schooley, R. T., Biswas, B., Gill, J. J., Hernandez-Morales, A., Lancaster, J.,
Lessor, L., et al. (2017). Development and use of personalized bacteriophage-
based therapeutic cocktails to treat a patient with a disseminated resistant
Acinetobacter baumannii infection. Antimicrob. Agents Chemother. 61:e0954-
17. doi: 10.1128/AAC.02221-18
Sulakvelidze, A., Alavidze, Z., and Morris, J. G. Jr. (2001). Bacteriophage therapy.
Antimicrob. Agents Chemother. 45, 649–659.
Suttle, C. A. (2013). Viruses: unlocking the greatest biodiversity on Earth. Genome
56, 542–544. doi: 10.1139/gen-2013-0152
Taylor, P. W., and Sommer, A. P. (2005). Towards rational treatment of bacterial
infections during extended space travel. Int. J. Antimicrob. Agents 26, 183–187.
doi: 10.1016/j.ijantimicag.2005.06.002
Thiebes, S., Schlesner, M., Brors, B., and Sunyaev, A. (2020). Distributed ledger
technology in genomics: a call for Europe. Eur. J. Hum. Genet. 28, 139–140.
doi: 10.1038/s41431-019-0512-4
Thiel, K. (2004). Old dogma, new tricks – 21st Century phage therapy. Nat.
Biotechnol. 2, 31–36. doi: 10.1038/nbt0104-31
Vandersteegen, K., Mattheus, W., Ceyssens, P. J., Bilocq, F., De Vos, D., Pirnay,
J.-P., et al. (2011). Microbiological and molecular assessment of bacteriophage
ISP for the control of Staphylococcus aureus. PLoS One 6:e24418. doi: 10.1371/
journal.pone.0024418
Verbeken, G., Pirnay, J. P., De Vos, D., Jennes, S., Zizi, M., Lavigne, R., et al. (2012).
Optimizing the European regulatory framework for sustainable bacteriophage
therapy in human medicine. Arch. Immunol. Ther. Exp. 60, 161–172. doi: 10.
1007/s00005-012-0175-0
Warwick, R. M., Chapman, J., Pruett, T. L., and Wang, H. (2013). Globally
consistent coding systems for medical products of human origin. Bull. World
Health Organ. 91, 314–314A. doi: 10.2471/BLT.12.116988
Yosef, I., Goren, M. G., Globus, R., Molshanski-Mor, S., and Qimron, U. (2017).
Extending the host range of bacteriophage particles for DNA transduction. Mol.
Cell. 66, 721.e3–728.e3. doi: 10.1016/j.molcel.2017.04.025
Conflict of Interest: The author declares that the research was conducted in the
absence of any commercial or financial relationships that could be construed as a
potential conflict of interest.
The handling editor declared past co-authorship with the author.
Copyright © 2020 Pirnay. This is an open-access article distributed under the terms
of the Creative Commons Attribution License (CC BY). The use, distribution or
reproduction in other forums is permitted, provided the original author(s) and the
copyright owner(s) are credited and that the original publication in this journal
is cited, in accordance with accepted academic practice. No use, distribution or
reproduction is permitted which does not comply with these terms.
Frontiers in Microbiology | www.frontiersin.org 8 June 2020 | Volume 11 | Article 1171
